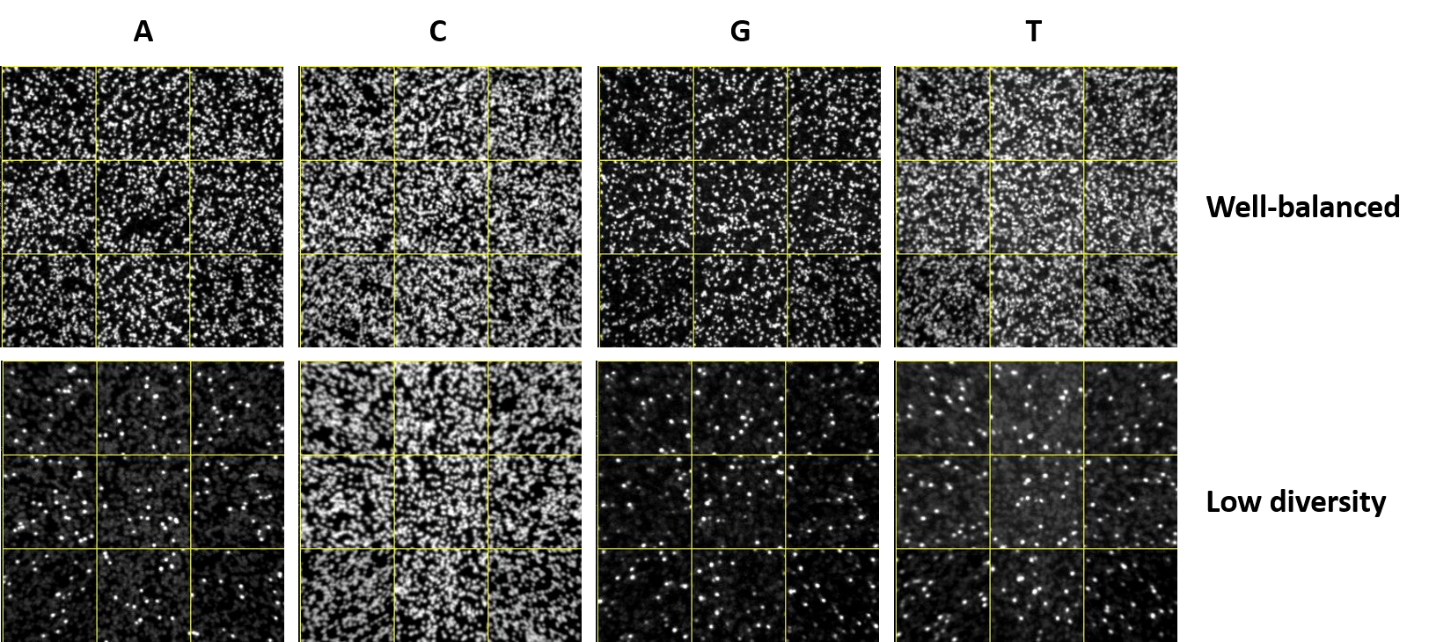
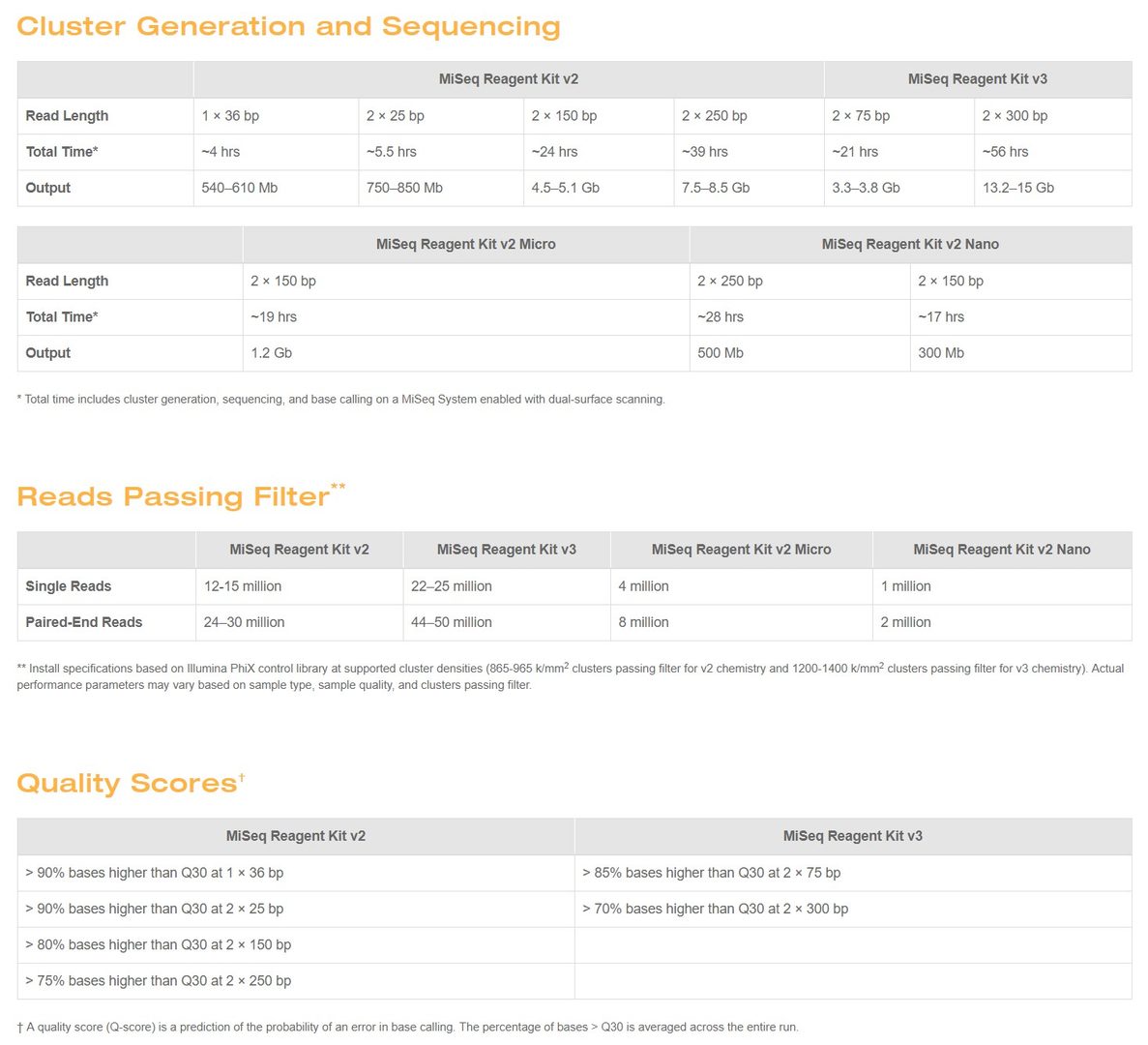
PLOS ONE: Large Scale Loss of Data in Low-Diversity Illumina Sequencing Libraries Can Be Recovered by Deferred Cluster Calling

gDNA Enrichment by a Transposase-based Technology for NGS Analysis of the Whole Sequence of BRCA1, BRCA2, and 9 Genes Involved in DNA Damage Repair | Protocol
Plotting % Occupied by % Pass Filter to optimize loading concentration for NovaSeq and iSeq 100 platforms

An adaptive decorrelation method removes Illumina DNA base-calling errors caused by crosstalk between adjacent clusters | Scientific Reports
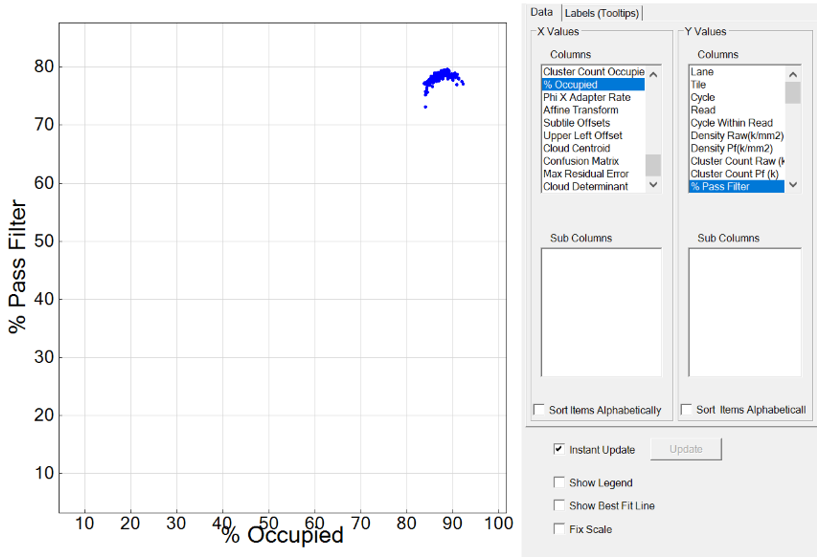
Plotting % Occupied by % Pass Filter to optimize loading concentration for NovaSeq and iSeq 100 platforms

Plotting % Occupied by % Pass Filter to optimize loading concentration for NovaSeq and iSeq 100 platforms
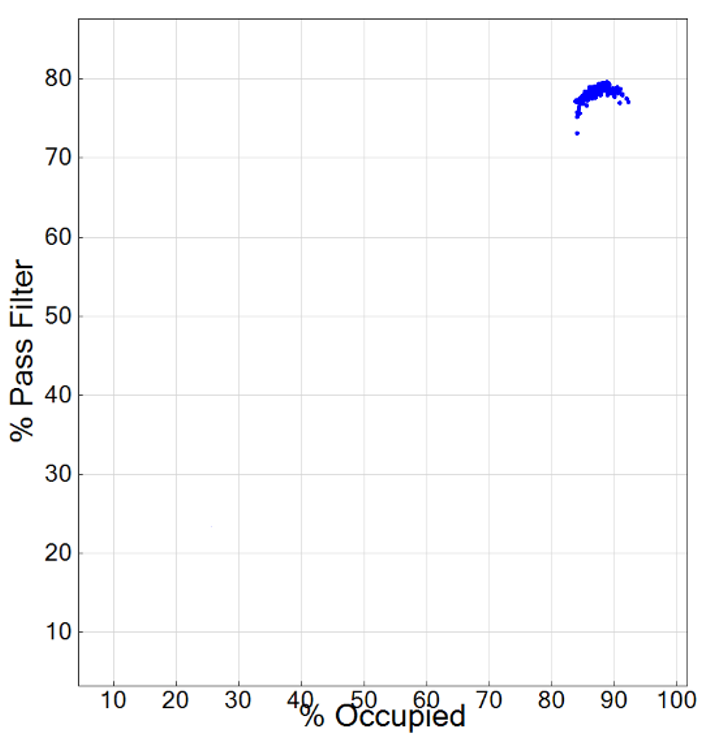
Plotting % Occupied by % Pass Filter to optimize loading concentration for NovaSeq and iSeq 100 platforms